Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
"label" - Generate Labels on Atoms
This section provides a tutorial on how to generate labels on atoms using the 'label' command in PyMol.
If you want to put some label on an atom, you can use the "label" command in the following syntax:
label atom_expression, string_express
The atom_expression can be a simple atom ID list like "id 1+2+4".
The string_express can be a Python string format pattern with atom properties as variables like 'id=%s, index=%s, name=%s" % (ID, index, name)'.
For example, the following commands label 3 atoms in a methane molecule:
PyMOL># load the carbon atom PyMOL>delete all PyMOL>load Carbon-Atom.sdf CmdLoad: loaded as "Carbon-Atom". PyMOL># add 4 hydrogen atoms to create methane PyMOL>h_add PyMOL># label 3 atoms PyMOL>label id 1+2+4, "id=%s, index=%s, name=%s" % (ID, index, name) Label: labelled 3 atoms.
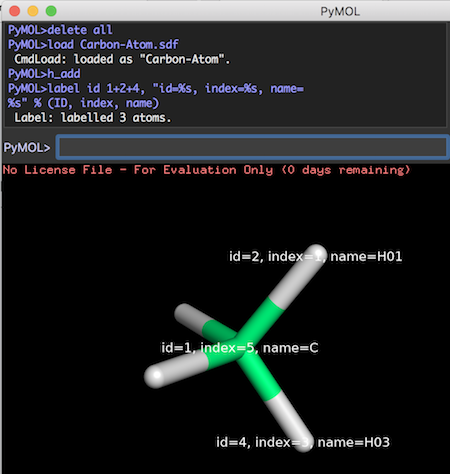
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
"get_extent" - Picked Atom Location
►"label" - Generate Labels on Atoms
Distance between Atoms in PyMol
Angle Formed by 3 Atoms in PyMol
Dihedral Angle Formed by 3 Atoms in PyMol
Use Selection Expressions in PyMol
"get_area" - Surface Areas of Atoms
Surface Area of Entire Molecule
"get_position" - Viewing Center
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)