Cheminformatics Tutorials - Herong's Tutorial Examples - v2.03, by Herong Yang
rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG - Molecule SVG Image
This section provides a quick introduction on the rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG class, which creates 2D molecule drawings in SVG format.
What Is rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG? - rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG class, together with its base class rdkit.Chem.Draw.rdMolDraw2D.MolDraw2D, provides functionalities to draw molecules in SVG vector image format.
Before playing with the rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG class, let's take a look at how to display SVG images on the screen.
The best way to display SVG image with Python is to use Jupyter Notebook, which uses the IPython kernel and related image display libraries as shown below.
IPython.display.SVG A class to create a SVG image object with given image data from a binary string, file, or URL.
- i = IPython.display.SVG(d) - Creates a SVG image object with given image data from a binary string.
- i = IPython.display.SVG(data=d) - Same as IPython.display.SVG(d).
- i = IPython.display.SVG(filename=f) - Creates a SVG image object with given image data from an external file.
- i = IPython.display.SVG(url=u) - Creates a SVG image object with given image data from a URL.
IPython.display.display(o) can also be used to display SVG images as described in the last tutorial.
Now, if we combine the power of RDKit and IPython, we can create and display a molecule SVG image very easily in Jupyter Notebook:
from rdkit import Chem
mol = Chem.MolFromSmiles('Cc1ccccc1')
from rdkit.Chem.Draw import rdMolDraw2D
d = rdMolDraw2D.MolDraw2DSVG(300, 300)
d.DrawMolecule(mol)
d.FinishDrawing()
p = d.GetDrawingText()
import IPython.display
i = IPython.display.SVG(p)
display(i)
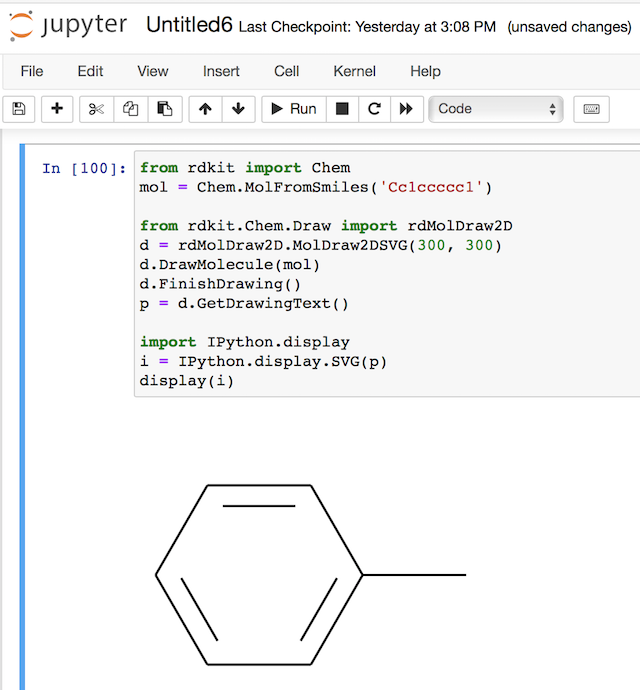
If you want to get a copy of the SVG string, you can call the print(p) method, as shown below:
>>> from rdkit import Chem
>>> mol = Chem.MolFromSmiles('Cc1ccccc1')
>>> from rdkit.Chem.Draw import rdMolDraw2D
>>> d = rdMolDraw2D.MolDraw2DSVG(300, 300)
>>> d.DrawMolecule(mol)
>>> d.FinishDrawing()
>>> p = d.GetDrawingText()
>>> print(p)
"<?xml version='1.0' encoding='iso-8859-1'?>
<svg version='1.1' baseProfile='full'
xmlns='http://www.w3.org/2000/svg'
xmlns:rdkit='http://www.rdkit.org/xml'
xmlns:xlink='http://www.w3.org/1999/xlink'
xml:space='preserve'
width='300px' height='300px' viewBox='0 0 300 300'>
<!-- END OF HEADER -->
<rect style='opacity:1.0;fill:#FFFFFF;stroke:none' width='300' height=...
<path class='bond-0 atom-0 atom-1' d='M 286.364,150 L 195.455,150' sty...
<path class='bond-1 atom-1 atom-2' d='M 195.455,150 L 150,228.73' styl...
<path class='bond-1 atom-1 atom-2' d='M 172.89,152.719 L 141.072,207.8...
<path class='bond-6 atom-6 atom-1' d='M 150,71.2704 L 195.455,150' sty...
<path class='bond-2 atom-2 atom-3' d='M 150,228.73 L 59.0909,228.73' s...
<path class='bond-3 atom-3 atom-4' d='M 59.0909,228.73 L 13.6364,150' ...
<path class='bond-3 atom-3 atom-4' d='M 68.0186,207.829 L 36.2005,152....
<path class='bond-4 atom-4 atom-5' d='M 13.6364,150 L 59.0909,71.2704'...
<path class='bond-5 atom-5 atom-6' d='M 59.0909,71.2704 L 150,71.2704'...
<path class='bond-5 atom-5 atom-6' d='M 72.7273,89.4522 L 136.364,89.4...
</svg>"
Table of Contents
SMILES (Simplified Molecular-Input Line-Entry System)
Open Babel: The Open Source Chemistry Toolbox
Using Open Babel Command: "obabel"
Generating SVG Pictures with Open Babel
Substructure Search with Open Babel
Similarity Search with Open Babel
Fingerprint Index for Fastsearch with Open Babel
Stereochemistry with Open Babel
Command Line Tools Provided by Open Babel
RDKit: Open-Source Cheminformatics Software
rdkit.Chem.rdchem - The Core Module
rdkit.Chem.rdmolfiles - Molecular File Module
rdkit.Chem.rdDepictor - Compute 2D Coordinates
►rdkit.Chem.Draw - Handle Molecule Images
What Is rdkit.Chem.Draw Module
MolToImage/MolToFile - Molecule PNG Image
rdkit.Chem.Draw.MolDrawing.DrawingOptions Class
rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DCairo - 2D Molecule Drawing
rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DCairo - Molecule PNG Image
►rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG - Molecule SVG Image
rdkit.Chem.Draw.rdMolDraw2D.MolDrawOptions - Drawing Options
Drawing Diagrams with MolDraw2DCairo and MolDraw2DSVG
Molecule Substructure Search with RDKit
rdkit.Chem.rdmolops - Molecule Operations
Daylight Fingerprint Generator in RDKit
Morgan Fingerprint Generator in RDKit
RDKit Performance on Substructure Search
Introduction to Molecular Fingerprints
OCSR (Optical Chemical Structure Recognition)
AlphaFold - Protein Structure Prediction