Cheminformatics Tutorials - Herong's Tutorial Examples - v2.03, by Herong Yang
Substructure Search with SMARTS Expressions
This section provides a tutorial example on how to use SMARTS expressions string to do substructure search using 'obabel ... -s ...' command with Open Babel.
What Are SMARTS Expressions? - A SMARTS expression is a sequence of atoms, bonds, and operators that can be evaluated to a substructure pattern. SMARTS expressions are divided into 3 types:
1. Atom Expressions - An expression that can be evaluated to an atom pattern using atom operations on atoms and atom expressions.
An atom expression must be enclosed in square bracktes, if it is not a single simple atom symbol.
Here are some examples of atom expressions of a single atom operand by itself. No operator is used. The object molecule must have at least one atom matching the given atom pattern.
Atom Description ---- ----------- C normal (aliphatic) carbon atom [#6] atom with 6 protons (carbon atom) c aromatic carbon atom [Ca] calcium atom [R] any atom on any ring [D3] any atom connected with 3 explicit bonds [X3] any atom connected with 3 total bonds (includes implicit H bonds) [h2] any atom connected with 2 implicit hydrogens [H2] any atom connected with 2 total hydrogens a any aromatic atom * any atom
Here are atom operators and their precedences:
Precedence Operation Description
---------- --------- -----------
4 [!e] NOT operation, negate expression e
The object molecule must have no atom matching
the given atom expression.
3 [e1&e2] AND operation, "e1" and "e2" are matched
The object molecule must have least one atom matching
both given atom expressions.
3 [e1e2] AND operation, same as above, & omitted
2 [e1,e2] OR operation, "e1" or "e2" is matched
The object molecule must have least one atom matching
one of the given atom expressions.
1 [e1;e2] AND operation with lower precedence,
Same as "&" operation, but evaluated after ",".
2. Bond Expressions - An expression that can be evaluated to a bond pattern using bond operations on bonds and bond expressions.
Here are some examples of bond expressions of one bond operand by itself. No operator is used. The object molecule must have at least one bond matching the given bond pattern.
Bond Description ---- ----------- - single bond (aliphatic), the default bond, can be omitted = double bond # triple bond / directional bond "up" \ directional bond "down"1 /? directional bond "up or unspecified" \? directional bond "down or unspecified" : aromatic bond @ any ring bond ~ any bond
Here are bond operators and their precedences:
Precedence Operation Description
---------- --------- -----------
4 !e NOT operation, negate expression e
The object molecule must have no bond matching
the given bond expression.
3 e1&e2 AND operation, "e1" and "e2" are matched
The object molecule must have least one bond matching
both given bond expressions.
3 e1e2 AND operation, same as above, & omitted
2 e1,e2 OR operation, take "e1" or "e2" is matched
The object molecule must have least one bond matching
one of the given bond expressions.
1 e1;e2 AND operation with lower precedence,
Same as "&" operation, but evaluated after ",".
3. Recursive SMARTS Expressions - An expression that can be evaluated to a SMARTS pattern using SMARTS operations on SMARTS and SMARTS expressions.
A SMARTS expression must be enclosed in bracktes and prefixed with "$", if it is used in a SMARTS operation.
Here are SMARTS operators and their precedences:
Precedence Operation Description
---------- --------- -----------
4 !$(e) NOT operation, negate expression e
The object molecule must not match the given
SMARTS expression.
3 $(e1)&$(e2) AND operation, "e1" and "e2" are matched
The object molecule must have least one bond matching
both given bond expressions.
3 $(e1)$(e2) AND operation, same as above, & omitted
2 $(e1),$(e2) OR operation, take "e1" or "e2" is matched
The object molecule must have least one bond matching
one of the given bond expressions.
1 $(e1);$(e2) AND operation with lower precedence,
Same as "&" operation, but evaluated after ",".
Open Babel Supports SMARTS expressions - You can use SMARTS expressions in the "-s ..." option in "obabel" commands to filter molecules that match given SMARTS expressions.
Here are some examples:
# C, C and O connected with single bonds herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s C-C-O c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # same as above with single bonds omitted herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s CCO c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # same as above atom expression with optional brackets added herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s C[C]O c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # double conditions on the middle atom herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s C[CH0]O c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # same as above with implicit & included herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s 'C[C&H0]O' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # bond expression used herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s 'C-,=O' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # same as above, but in a native way herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s 'C!#O' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # bad bond expression, no bond can be both single and double. herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s 'C-=O' 0 molecules converted # poor bond expression, a single bond is also an any bond. herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s 'C-~O' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # matching aromatic C and connected with 1 H herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s '[c;H1]' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # matching aromatic C and connected with 0 H herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles -s '[c;H0]' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted # nested SMARTS expressions herong$ obabel "-:c1cc(ccc1CC(C(=O)O)N)O" -o smiles \ -s '[C;H0]-,=[$([O;H1]),$([O;H0])]' c1cc(ccc1CC(C(=O)O)N)O 1 molecule converted
You can validate the above matching result by looking at the tyrosine molecule structure below:
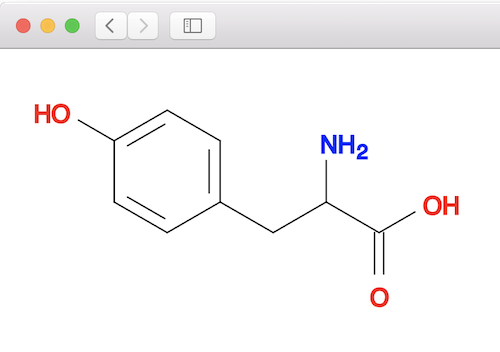
Table of Contents
SMILES (Simplified Molecular-Input Line-Entry System)
Open Babel: The Open Source Chemistry Toolbox
Using Open Babel Command: "obabel"
Generating SVG Pictures with Open Babel
►Substructure Search with Open Babel
"obabel -s ..." Command - Substructure Search
Substructure Search with Wildcard Atom "*"
Substructure Search with Wildcard Bond "~"
►Substructure Search with SMARTS Expressions
Similarity Search with Open Babel
Fingerprint Index for Fastsearch with Open Babel
Stereochemistry with Open Babel
Command Line Tools Provided by Open Babel
RDKit: Open-Source Cheminformatics Software
rdkit.Chem.rdchem - The Core Module
rdkit.Chem.rdmolfiles - Molecular File Module
rdkit.Chem.rdDepictor - Compute 2D Coordinates
rdkit.Chem.Draw - Handle Molecule Images
Molecule Substructure Search with RDKit
rdkit.Chem.rdmolops - Molecule Operations
Daylight Fingerprint Generator in RDKit
Morgan Fingerprint Generator in RDKit
RDKit Performance on Substructure Search
Introduction to Molecular Fingerprints
OCSR (Optical Chemical Structure Recognition)
AlphaFold - Protein Structure Prediction