Cheminformatics Tutorials - Herong's Tutorial Examples - v2.03, by Herong Yang
Change Display Command on Open Babel GUI
This section provides a tutorial example on how to change structure display command on Open Babel GUI to use Google Chrome browser.
By default, Open Babel GUI uses Firefox to display molecule structure. If you don't have Firefox installed, you will get an error:
Obgui Error: Execution of command "firefox ..." failed. System cannot find the file.
To fix the problem, you can change the display command as shown in this tutorial.
1. Start Open Babel GUI and go to "View > Configure Structure Display" menu. You see a small window showing the current display command.
2. Enter a new command to use a different browser. For example, enter the following to use the Google Chrome browser:
\Progra~1\Google\Chrome\Application\Chrome file:/// C:\Users\herong\AppData\Local\Temp/gui.svg
3. Enter a new molecule in the input section on the left.
4. Click to select the "Display in ..." checkbox in the output section on the right.
5. Click the "CONVERT" button. You see the molecule structure displayed in the Google Chrome browser.
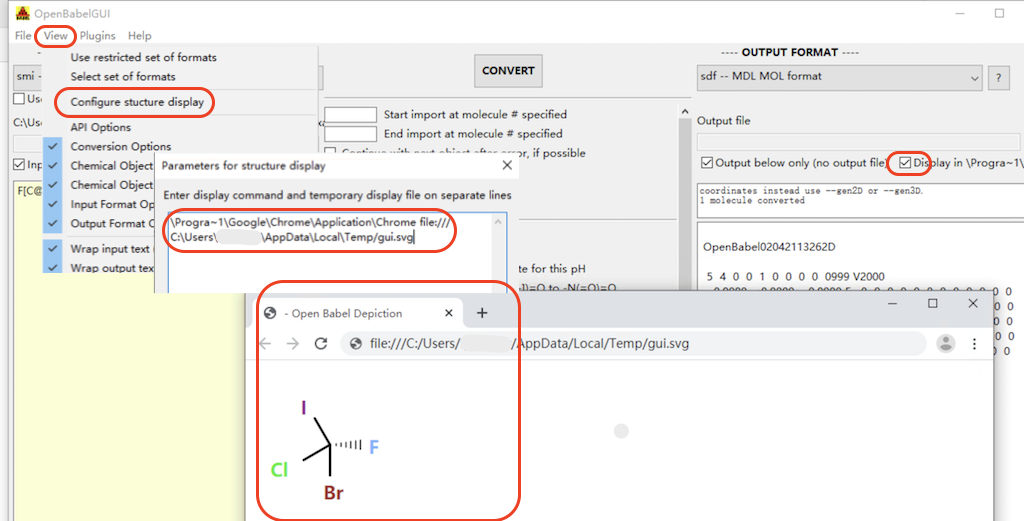
Table of Contents
SMILES (Simplified Molecular-Input Line-Entry System)
►Open Babel: The Open Source Chemistry Toolbox
Install Open Babel with Anaconda
Install Open Babel on Windows Computers
Run Open Babel GUI on Windows Computers
►Change Display Command on Open Babel GUI
Open Babel Installation Options on Linux
Install Open Babel Binary Package on CentOS
"Open Babel Error in LoadAllPlugins" Error
Install Open Babel from Source Code
Install Open Babel 2.4.1 from Source Code
Open Babel Installation Options on macOS
Install Open Babel Binary Package on macOS
Using Open Babel Command: "obabel"
Generating SVG Pictures with Open Babel
Substructure Search with Open Babel
Similarity Search with Open Babel
Fingerprint Index for Fastsearch with Open Babel
Stereochemistry with Open Babel
Command Line Tools Provided by Open Babel
RDKit: Open-Source Cheminformatics Software
rdkit.Chem.rdchem - The Core Module
rdkit.Chem.rdmolfiles - Molecular File Module
rdkit.Chem.rdDepictor - Compute 2D Coordinates
rdkit.Chem.Draw - Handle Molecule Images
Molecule Substructure Search with RDKit
rdkit.Chem.rdmolops - Molecule Operations
Daylight Fingerprint Generator in RDKit
Morgan Fingerprint Generator in RDKit
RDKit Performance on Substructure Search
Introduction to Molecular Fingerprints
OCSR (Optical Chemical Structure Recognition)
AlphaFold - Protein Structure Prediction