Cheminformatics Tutorials - Herong's Tutorial Examples - v2.01, by Herong Yang
rdkit.Chem.Draw.MolDrawing.DrawingOptions Class
This section provides a tutorial example on how to use rdkit.Chem.Draw.DrawingOptions class, presenting a set of molecule drawing options. But I could not find a way to use it.
What Is rdkit.Chem.Draw.MolDrawing.DrawingOptions Class? - rdkit.Chem.Draw.MolDrawing.DrawingOptions class prepresents a set of molecule drawing options. You can use them to control the appearance of molecule images.
rdkit.Chem.Draw.MolDrawing.DrawingOptions contains the following attributes as molecule drawing options:
atomLabelDeuteriumTritium = False
atomLabelFontFace = 'sans'
atomLabelFontSize = 12
atomLabelMinFontSize = 7
atomNumberOffset = 0
bgColor = (1, 1, 1)
bondLineWidth = 1.2
colorBonds = True
coordScale = 1.0
dash = (4, 4)
dblBondLengthFrac = 0.8
dblBondOffset = 0.25
defaultColor = (1, 0, 0)
dotsPerAngstrom = 30
elemDict = {0: (0.5, 0.5, 0.5), 1: (0.55, 0.55, 0.55), 7: (0, 0, 1), ...}
includeAtomNumbers = False
noCarbonSymbols = True
radicalSymbol = '∙'
selectColor = (1, 0, 0)
showUnknownDoubleBonds = True
useFraction = 0.85
wedgeDashedBonds = True
Here is an example session of using the rdkit.Chem.Draw.MolDrawing.DrawingOptions class in a Jupyter Notebook environment.
from rdkit import Chem
m = Chem.MolFromSmiles('Cc1ccccc1')
from rdkit.Chem.Draw.MolDrawing import DrawingOptions
opts = DrawingOptions()
opts.includeAtomNumbers = True
opts.bondLineWidth = 4
from rdkit.Chem import Draw
p = Draw.MolToImage(m, options=opts)
p
Unfortunately, this example is not working as I expected. The DrawingOptions object seems be ignored by the MolToImage() method. I am getting the same molecule image as with default options. See the Notebook below:
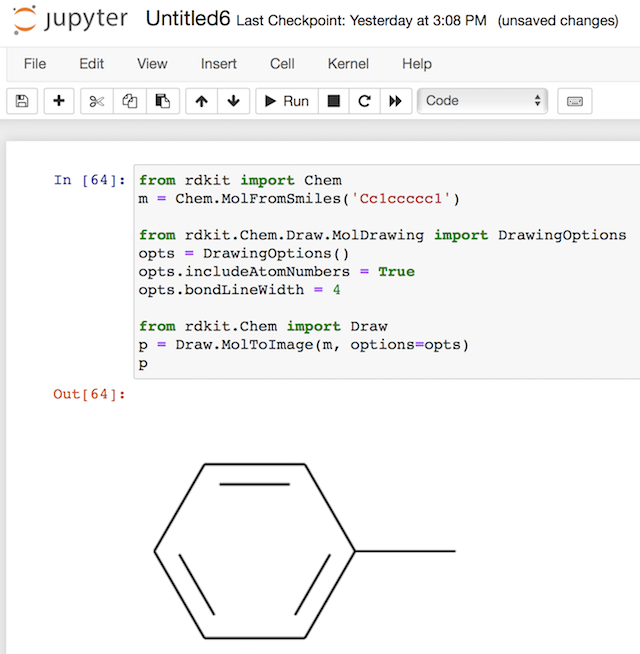
Table of Contents
SMILES (Simplified Molecular-Input Line-Entry System)
Open Babel: The Open Source Chemistry Toolbox
Using Open Babel Command: "obabel"
Generating SVG Pictures with Open Babel
Substructure Search with Open Babel
Similarity Search with Open Babel
Fingerprint Index for Fastsearch with Open Babel
Stereochemistry with Open Babel
Command Line Tools Provided by Open Babel
RDKit: Open-Source Cheminformatics Software
rdkit.Chem.rdchem - The Core Module
rdkit.Chem.rdmolfiles - Molecular File Module
rdkit.Chem.rdDepictor - Compute 2D Coordinates
►rdkit.Chem.Draw - Handle Molecule Images
What Is rdkit.Chem.Draw Module
MolToImage/MolToFile - Molecule PNG Image
►rdkit.Chem.Draw.MolDrawing.DrawingOptions Class
rdkirdkit.Chem.Draw.rdMolDraw2D.MolDraw2DCairo - 2D Molecule Drawing
rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DCairo - Molecule PNG Image
rdkit.Chem.Draw.rdMolDraw2D.MolDraw2DSVG - Molecule SVG Image
rdkit.Chem.Draw.rdMolDraw2D.MolDrawOptions - Drawing Options
Drawing Diagrams with MolDraw2DCairo and MolDraw2DSVG
Molecule Substructure Search with RDKit
rdkit.Chem.rdmolops - Molecule Operations
Daylight Fingerprint Generator in RDKit
Morgan Fingerprint Generator in RDKit
RDKit Performance on Substructure Search
Introduction to Molecular Fingerprints
OCSR (Optical Chemical Structure Recognition)
AlphaFold - Protein Structure Prediction